Non-rigid Registration¶
Non-rigid registration methods are capable of aligning images where correspondence cannot be achieved without localized deformations and can therefore better accomodate anatomical, physiological and pathological variability between patients.
B-splines are often used to parameterize a free-form deformation (FFD) field. This is a much harder registration problem than any of the previous examples due to a much higher-dimensional parameter space and we are therefore best off using a multi-resolution approach with affine initialization. This is very easy to do in SimpleElastix.
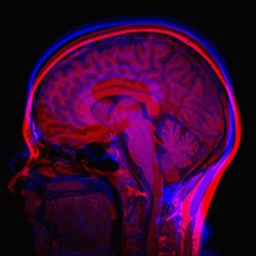
Consider the following mean image of two different subjects.
The following code runs multi-resolution affine initialization and starts a non-rigid method multi-resolution non-rigid method using the affine transform as initialization :
import SimpleITK as sitk
elastixImageFilter = sitk.ElastixImageFilter()
elastixImageFilter.SetFixedImage(sitk.ReadImage("fixedImage.nii"))
elastixImageFilter.SetMovingImage(sitk.ReadImage("movingImage.nii"))
parameterMapVector = sitk.VectorOfParameterMap()
parameterMapVector.append(sitk.GetDefaultParameterMap("affine"))
parameterMapVector.append(sitk.GetDefaultParameterMap("bspline"))
elastixImageFilter.SetParameterMap(parameterMapVector)
elastixImageFilter.Execute()
sitk.WriteImage(elastixImageFilter.GetResultImage())
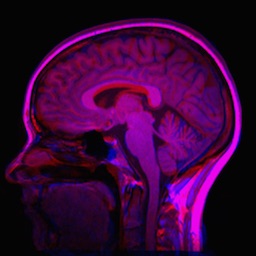
The result image is seen below.
In this case, we are able to compensate for many non-rigid differences between the two images. Note, however, that brain image registration is a difficult to task because of complex anatomical variations. Entire registration packages are dedicated to brain image processing. You might want to consider a more refined approach in critical applications.
In the next section we introduce groupwise registration, where many images are registered simultaneously a mean frame of reference.